Topic overview
Molecular Microbiology
Click to learn more and explore related projects.
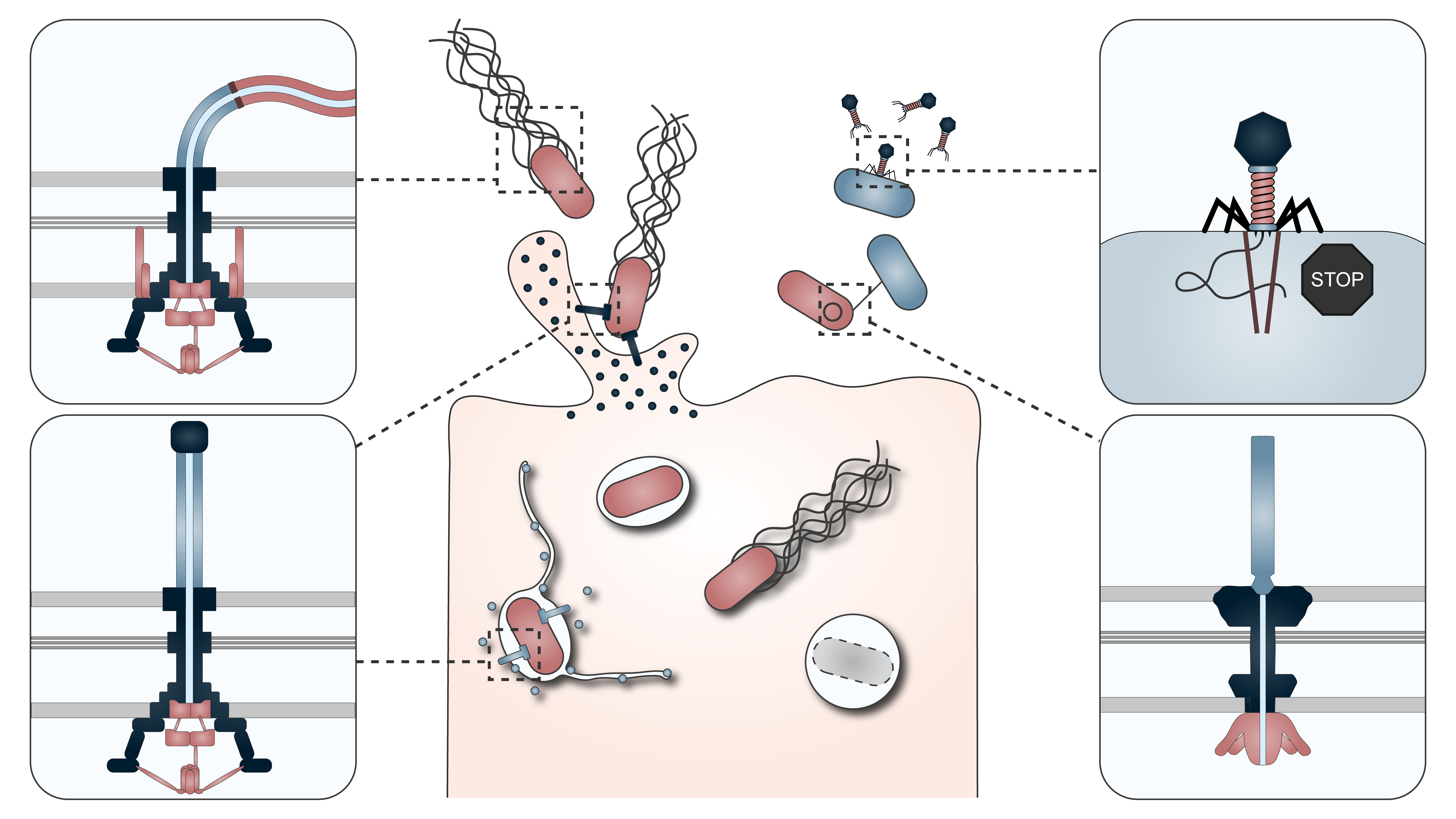 Flagella
vT3SS
Host‑pathogen interaction
Phage defense
T4SS
Development of advanced image analysis tools
Flagella
vT3SS
Host‑pathogen interaction
Phage defense
T4SS
Development of advanced image analysis tools
Funding sources
Molecular Microbiology